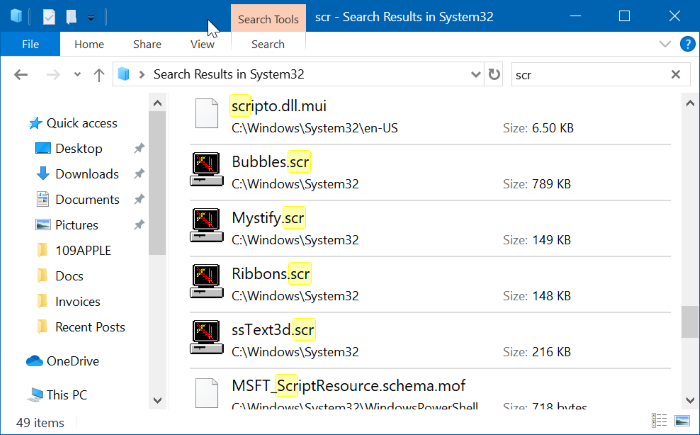
Note: Additional features are available in SnapGene. SnapGene Viewer is a revolutionary software app that allows molecular biologists to create, browse, and share richly annotated DNA sequence files up to 1 Gbp in length.īeautiful maps Zooming for chromosomes Primer binding sites Enzyme sites and properties Sequence trace viewer Restriction site overview Clear methylation indicators Reading frames for gene fusions Feature and primer lists Versatile file conversion Automatic feature annotation This is the Free version, because there should be no barriers to seeing your data. SnapGene Viewer includes the same rich visualization, annotation, and sharing capabilities as the fully enabled SnapGene software. Requirements: Windows XP / Vista / Windows 7 / Windows 8 / Windows 10 / Windows XP64 / Vista64 / Windows 7 64 / Windows 8 64 / Windows 10 64Īuthor / Product: GSL Biotech LLC / SnapGene Viewer On the webpage of pcDNA4/TO click the pcDNA4 TO.dna link to save the file: Annotation: every possible piece of information about a sequenceĭownload the annotated sequence of pcDNA4/TO from SnapGene.SnapGene Viewer 3.2.1 Download for Windows 10, 8, 7 Open the annotated sequence of pcDNA4/TO in SnapGene. You now see the circular view of the vector with restriction sites automatically visualized:Įxercise: load the sequence of vector pAcGFP1-C1 Open SnapGene and select Open in the startup menu:īrowse to the pcDNA4 TO.dna file and open it. This vector is available in SnapGene's list of fluorescent protein genes and plasmids. On the webpage of pAcGFP1-C1 click pAcGFP1-C1.dna: Try to load it without peeking at the solution. Loading vector sequences from other resources Go back to SnapGene and click the Open button in the top menu:īrowse to the pAcGFP1-C1.dna file and open it. However, not all vector sequences are stored in the SnapGene database. Fortunately, many vectors are available from Addgene, an organization that improves sharing of plasmids. In SnapGene you can load sequences directly from Genbank or you can load them by importing a file. When you make a post, and it does not appear, it went into moderation. Some posts are auto-moderated to reduce spam, including links and swear words. Even if you don't load them from Genbank it is still a good idea to import them in Genbank format (and not in Fasta format). Comment Rules & Etiquette - We welcome all comments from our readers, but any comment section requires some moderation. Genbank format not only contains a sequence but also all annotations. 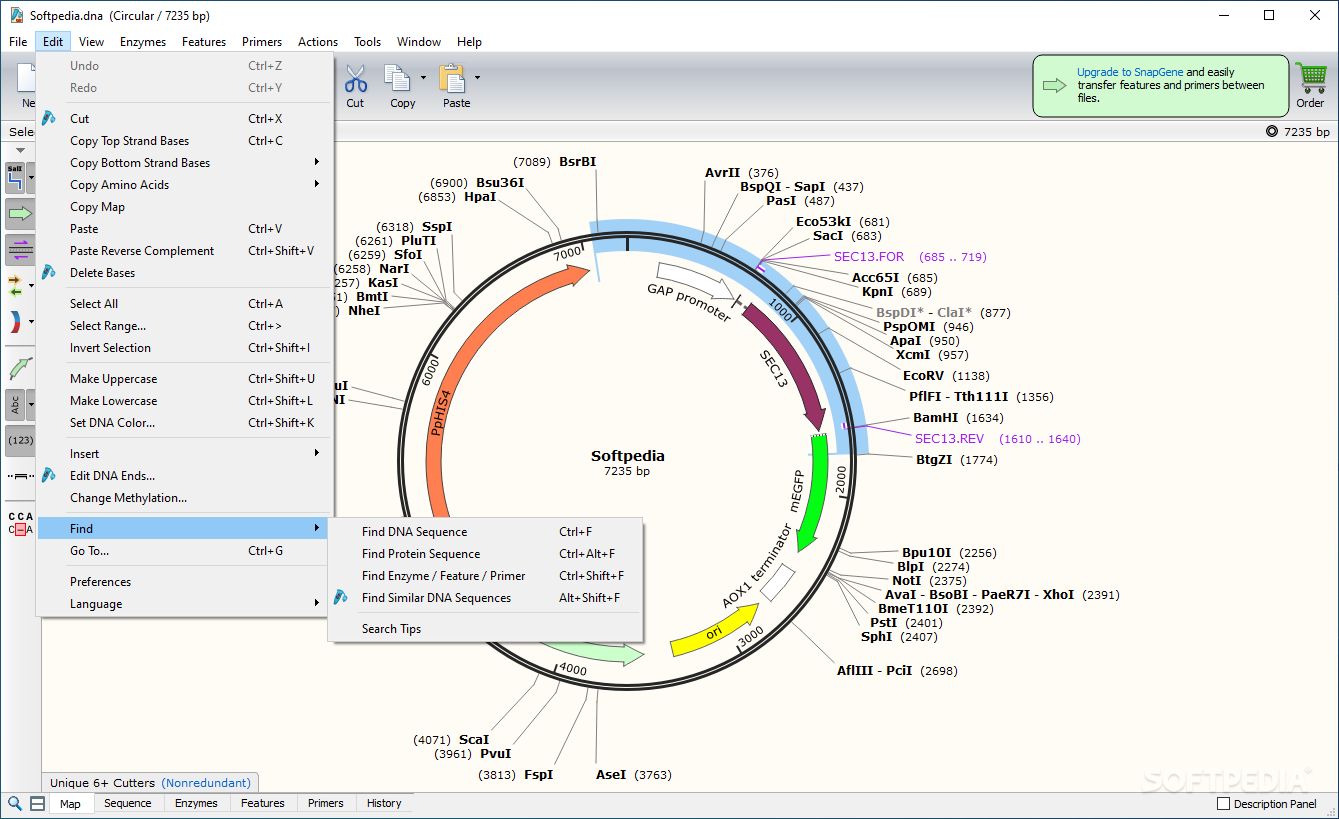
If you load the annotations (in Genbank format) SnapGene will automatically show them. On the next page containing the full sequence of the vector, click the Analyze Sequence button on the right: On the webpage of mRFP-Rab5 go to the SEQUENCE INFORMATION section and click Full (1) next to Depositor Sequences: Obtain the annotated sequence of mRFP-Rab5 from Addgenes. On the next page select the Sequence tab: #Snapgene viewer version full To include the annotation, you need the sequence in Genbank format:Ĭopy the text and paste it in Notepad. 
Open the annotated sequence of mRFP-Rab5 in SnapGene.Ĭlick the Open button in the top menu, browse to the mRFPrab5.txt file and open it. Since it's a vector sequence, select circular:Ĭlick OK to open the sequence in SnapGene.Ĭhanging annotation of sequences in SnapGeneĪlthough we imported mRFP1-Rab5 in Genbank format, the annotation that was included in the file was not complete: SnapGene will ask you wether the sequence is linear or circular.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
